|
Information for users of fluvial software about new releases and developments. Flumen v3.0 releasedFlumen has been optimised for higher performance and dealing with large meshes. The main new options of Flumen v3.0 are: dynamic storage allocation: allows large models (> 10 M cells, depending on RAM)
support of multi-core
CPU. The number of threads to be used is defined in the input
file: less storage space (.res file). Version 3.0 can be downloaded from here. Simp 3.0 releasedThe new version of Simp has been optimized for handling large data sets. For this reason the data is divided into tiles (computational units) that are treated one by one. This is not only much faster, it also uses less storage and therefore allows to apply finer grids for the filtering algorithm. The default value for the raster width is now reduced to 0.5 m (before 1.0 m). Tips & Tricks --> OptionsOften many simulation runs have to be made with only small differences in the input parameters. In the end there are multiple input files that differ only by one or two lines. Using “options” things can be handled much more comfortable. In the input file you can write e.g. {optionA
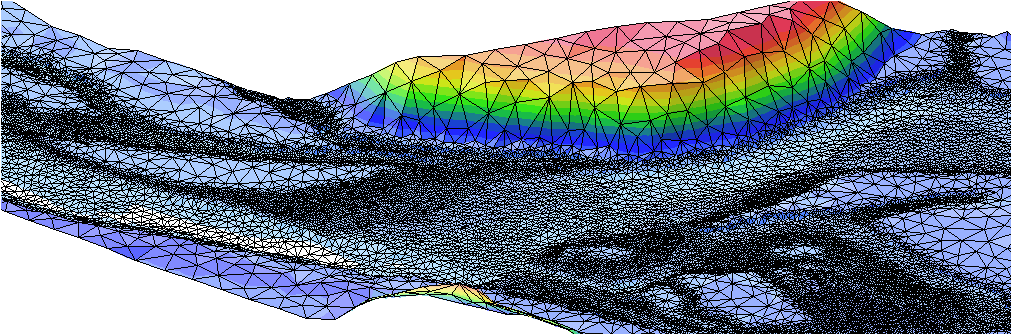
inflow 150. Then when starting the program, in the command shell you write flumen input_file @optionA to get an inflow of 150 m3/s in the simulation. Using options is very useful for managing input files (you need only one!) and thus avoiding modeling errors. More details of how to use options are explained here. Next IssueA new module has been added to the preprocessor FLUVIZ to produce efficient meshes regarding the CFL limitation. Together with TRIANGLE quality meshes are created with variable cell sizes (small areas in shallow regions, see figure below).
The new feature will be presented in more detail in the next issue. |
|
Impressum
/ contact |
|
For comments or to subscribe/unsubscribe from the mailing list, please send e-mail to info@fluvial.ch |